cppFunction('
double f1(NumericVector x) {
int n = x.size();
double y = 0;
for(int i = 0; i < n; ++i) {
y += x[i] / n;
}
return y;
}
')
cppFunction('
NumericVector f2(NumericVector x) {
int n = x.size();
NumericVector out(n);
out[0] = x[0];
for(int i = 1; i < n; ++i) {
out[i] = out[i - 1] + x[i];
}
return out;
}
')
cppFunction('
bool f3(LogicalVector x) {
int n = x.size();
bool any = false;
for(int i = 0; i < n; ++i) {
if (x[i]) any = true;
}
return any;
}
')
cppFunction('
int f4(Function pred, List x) {
int n = x.size();
for(int i = 0; i < n; ++i) {
LogicalVector res = pred(x[i]);
if (res[0]) return i + 1;
}
return 0;
}
')
cppFunction('
NumericVector f5(NumericVector x, NumericVector y) {
int n = std::max(x.size(), y.size());
NumericVector x1 = rep_len(x, n);
NumericVector y1 = rep_len(y, n);
NumericVector out(n);
for (int i = 0; i < n; ++i) {
out[i] = std::min(x1[i], y1[i]);
}
return out;
}
')Solutions to Exercises Week 4
Exercise 25.2.6.1
Question: With the basics of C++ in hand, it’s now a great time to practice by reading and writing some simple C++ functions. For each of the following functions, read the code and figure out what the corresponding base R function is. You might not understand every part of the code yet, but you should be able to figure out the basics of what the function does.
A: We can try running each function and seeing what they do. For example, we can run the following code:
x <- 1:4
f1(x) - mean(x) # f1 is the mean[1] 0f2(x) - cumsum(x) # f2 is the cumsum[1] 0 0 0 0f3(c(F, F, T, F)) == any(c(F, F, T, F)) # f3 is the any function[1] TRUEf4(function(x) c(x > 3, x > 6), 1:5) # finds the first in the list where the function has a truthy value[1] 4f5(1:5, 6:10) == apply(rbind(1:5, 6:10), MARGIN = 2, min) # f5 applies the min function to each column of the matrix[1] TRUE TRUE TRUE TRUE TRUEExercise 25.2.6.2
To practice your function writing skills, convert the following functions into C++. For now, assume the inputs have no missing values. + all(). + cumprod(), cummin(), cummax(). + diff(). Start by assuming lag 1, and then generalise for lag n. + range(). + var(). Read about the approaches you can take on Wikipedia. Whenever implementing a numerical algorithm, it’s always good to check what is already known about the problem.
cppFunction('
bool all_cpp(LogicalVector x) {
int n = x.size();
for(int i = 0; i < n; ++i) {
if (!x[i]) return false;
}
return true;
}
')
cppFunction('
double cumprod_cpp(NumericVector x) {
int n = x.size();
double prod = 1;
for(int i = 0; i < n; ++i) {
prod *= x[i];
}
return prod;
}
')
cppFunction('
NumericVector cummin_cpp(NumericVector x) {
int n = x.size();
double min = x[0];
NumericVector out(n);
out[0] = min;
for(int i = 1; i < n; ++i) { // this is bad, why?
if (x[i] < min) min = x[i];
out[i] = min;
}
return out;
}
')
cppFunction('
NumericVector diff_cpp(NumericVector x, int lag = 1) {
int n = x.size();
NumericVector out(n - lag);
for(int i = 0; i < n - lag; ++i) {
out[i] = x[i + lag] - x[i];
}
return out;
}
')
cppFunction('
NumericVector range_cpp(NumericVector x) {
double min = x[0], max = x[0];
int n = x.size();
for(int i = 1; i < n; ++i) {
if (x[i] < min) min = x[i];
if (x[i] > max) max = x[i];
}
NumericVector out = NumericVector::create(min, max);
return out;
}
')
cppFunction('
double var_cpp(NumericVector x) {
int n = x.size();
double sum = 0, sumsq = 0;
for(int i = 0; i < n; ++i) {
sum += x[i];
sumsq += x[i] * x[i];
}
return (sumsq - sum * sum / n) / (n - 1);
}
')Question: See attached Rcpp file
all_cpp(c(F, T, T, T))[1] FALSEcumprod_cpp(1:5)[1] 120cummin_cpp(c(1, 2, 3, 0, 1))[1] 1 1 1 0 0diff_cpp(1:10, lag = 1)[1] 1 1 1 1 1 1 1 1 1diff_cpp(1:10, lag = 2)[1] 2 2 2 2 2 2 2 2range_cpp(1:10)[1] 1 10var_cpp(rnorm(1e5))[1] 1.003504Exercise 25.4.5.1
Question: Rewrite any of the functions from the first exercise of Section 25.2.6 to deal with missing values. If na.rm is true, ignore the missing values. If na.rm is false, return a missing value if the input contains any missing values. Some good functions to practice with are min(), max(), range(), mean(), and var().
Answer: Use the decorator design pattern,
.na.rm_decorator <- function(fun) {
function(x, na.rm = FALSE, ...) {
if (na.rm) {
x <- x[!is.na(x)]
} else if (any(is.na(x))) {
return(NA)
}
return(fun(x, ...))
}
}
## shorter version using only one if statement
.na.rm_decorator_2 <- function(fun) {
function(x, na.rm = FALSE, ...) {
if (!na.rm && any(is.na(x))) {
return(NA)
}
x <- x[!is.na(x)]
return(fun(x, ...))
}
}Use our decorators
mean_decorated <- .na.rm_decorator(f1)
mean_decorated(c(1, 2, NA, 3, 4))[1] NAmean_decorated(c(1, 2, NA, 3, 4), na.rm = TRUE)[1] 2.5mean_decorated(c(1, 2, 3, 4))[1] 2.5diff_decorated <- .na.rm_decorator_2(diff_cpp)
diff_decorated(c(1, 2, NA, 3, 4), lag = 2)[1] NAdiff_decorated(c(1, 2, NA, 3, 4), na.rm = TRUE, lag = 2)[1] 2 2diff_decorated(c(1, 2, 3, 4), lag = 2)[1] 2 2Exercise 25.4.5.2
Question: Rewrite cumsum() and diff() so they can handle missing values. Note that these functions have slightly more complicated behaviour.
Answer: Use the decorator pattern again.
# For cumsum: use the first decorator (treats NAs as 0 when na.rm = TRUE)
cumsum_decorated <- .na.rm_decorator(f2)
# For diff: use the second decorator (removes NAs when na.rm = TRUE)
diff_decorated <- .na.rm_decorator_2(diff_cpp)
# Test the decorated functions
test_vec <- c(1, 2, NA, 4, 5)
# Test cumsum
cumsum_decorated(test_vec, na.rm = FALSE) # Should return NAs[1] NAcumsum_decorated(test_vec, na.rm = TRUE) # Should treat NA as 0[1] 1 3 7 12# Test diff
diff_decorated(test_vec, lag = 1, na.rm = FALSE) # Should return NAs[1] NAdiff_decorated(test_vec, lag = 1, na.rm = TRUE) # Should work on non-NA values[1] 1 2 1Bonus Example: Currying with Kernel Density Estimation (AI generated)
Question: How can we use currying (function composition and partial application) to create flexible and reusable kernel density estimation functions?
A: In U2.qmd, we implemented kernel density estimation with the Epanechnikov kernel. Now let’s see how currying can make our code more modular and elegant.
Step 1: Define Different Kernel Functions
First, let’s define several kernel functions:
# Epanechnikov kernel (from U2.qmd)
epanechnikov_kernel <- function(z) (abs(z) <= 1) * (1 - z^2) * 3 / 4
# Gaussian kernel
gaussian_kernel <- function(z) (1 / sqrt(2 * pi)) * exp(-0.5 * z^2)
# Uniform kernel
uniform_kernel <- function(z) (abs(z) <= 1) * 0.5
# Triangular kernel
triangular_kernel <- function(z) (abs(z) <= 1) * (1 - abs(z))Step 2: Create a General Kernel Density Function
Now let’s create a general function that can work with any kernel:
# General kernel density estimation function
kern_dens_general <- function(x, h, kernel_func, n = 512) {
rg <- range(x)
xx <- seq(rg[1] - 3 * h, rg[2] + 3 * h, length.out = n)
y <- numeric(n)
for (i in seq_along(xx)) {
y[i] <- mean(kernel_func((xx[i] - x) / h)) / h
}
list(x = xx, y = y)
}Step 3: Using Currying to Create Specialized Functions
Now comes the magic of currying! We can create specialized functions by “currying” our general function:
# Curry by kernel type - creates functions that use specific kernels
epanechnikov_kde <- function(x, h, n = 512) kern_dens_general(x, h, epanechnikov_kernel, n)
gaussian_kde <- function(x, h, n = 512) kern_dens_general(x, h, gaussian_kernel, n)
uniform_kde <- function(x, h, n = 512) kern_dens_general(x, h, uniform_kernel, n)
# Curry by bandwidth - creates functions with fixed bandwidths
epanechnikov_bw_half <- function(x, n = 512) epanechnikov_kde(x, h = 0.5, n)
epanechnikov_bw_one <- function(x, n = 512) epanechnikov_kde(x, h = 1.0, n)
epanechnikov_bw_two <- function(x, n = 512) epanechnikov_kde(x, h = 2.0, n)Step 4: Advanced Currying - Function Factories
We can create “function factories” that generate curried functions:
# Factory function that creates kernel-specific KDE functions
make_kernel_kde <- function(kernel_func) {
function(x, h, n = 512) kern_dens_general(x, h, kernel_func, n)
}
# Factory function that creates bandwidth-specific functions
make_bandwidth_kde <- function(kde_func, bandwidth) {
function(x, n = 512) kde_func(x, h = bandwidth, n)
}
# Create kernel-specific functions using the factory
epa_kde <- make_kernel_kde(epanechnikov_kernel)
gauss_kde <- make_kernel_kde(gaussian_kernel)
# Create bandwidth-specific functions
epa_narrow <- make_bandwidth_kde(epa_kde, 0.3)
epa_medium <- make_bandwidth_kde(epa_kde, 0.8)
epa_wide <- make_bandwidth_kde(epa_kde, 1.5)Step 5: Demonstration with Data
Let’s test our curried functions with the infrared data:
# Load the data (same as U2.qmd)
infrared <- read.table(here::here("data", "infrared.txt"), header = TRUE)
F12 <- infrared$F12
log_F12 <- log(F12)
# Test our curried functions
dens_narrow <- epa_narrow(log_F12)
dens_medium <- epa_medium(log_F12)
dens_wide <- epa_wide(log_F12)
# Compare different kernels with same bandwidth
dens_epa <- epa_kde(log_F12, h = 0.8)
dens_gauss <- gauss_kde(log_F12, h = 0.8)
dens_uniform <- make_kernel_kde(uniform_kernel)(log_F12, h = 0.8)Step 6: Visualization
library(ggplot2)
# Create data frames for plotting
df_bandwidths <- data.frame(
x = rep(dens_narrow$x, 3),
y = c(dens_narrow$y, dens_medium$y, dens_wide$y),
bandwidth = rep(c("Narrow (0.3)", "Medium (0.8)", "Wide (1.5)"), each = length(dens_narrow$x))
)
df_kernels <- data.frame(
x = rep(dens_epa$x, 3),
y = c(dens_epa$y, dens_gauss$y, dens_uniform$y),
kernel = rep(c("Epanechnikov", "Gaussian", "Uniform"), each = length(dens_epa$x))
)
# Plot different bandwidths
p1 <- ggplot(df_bandwidths, aes(x = x, y = y, color = bandwidth)) +
geom_line(size = 1) +
labs(title = "Effect of Bandwidth (Epanechnikov Kernel)",
x = "log(F12)", y = "Density", color = "Bandwidth") +
theme_minimal()
# Plot different kernels
p2 <- ggplot(df_kernels, aes(x = x, y = y, color = kernel)) +
geom_line(size = 1) +
labs(title = "Different Kernels (h = 0.8)",
x = "log(F12)", y = "Density", color = "Kernel") +
theme_minimal()
print(p1)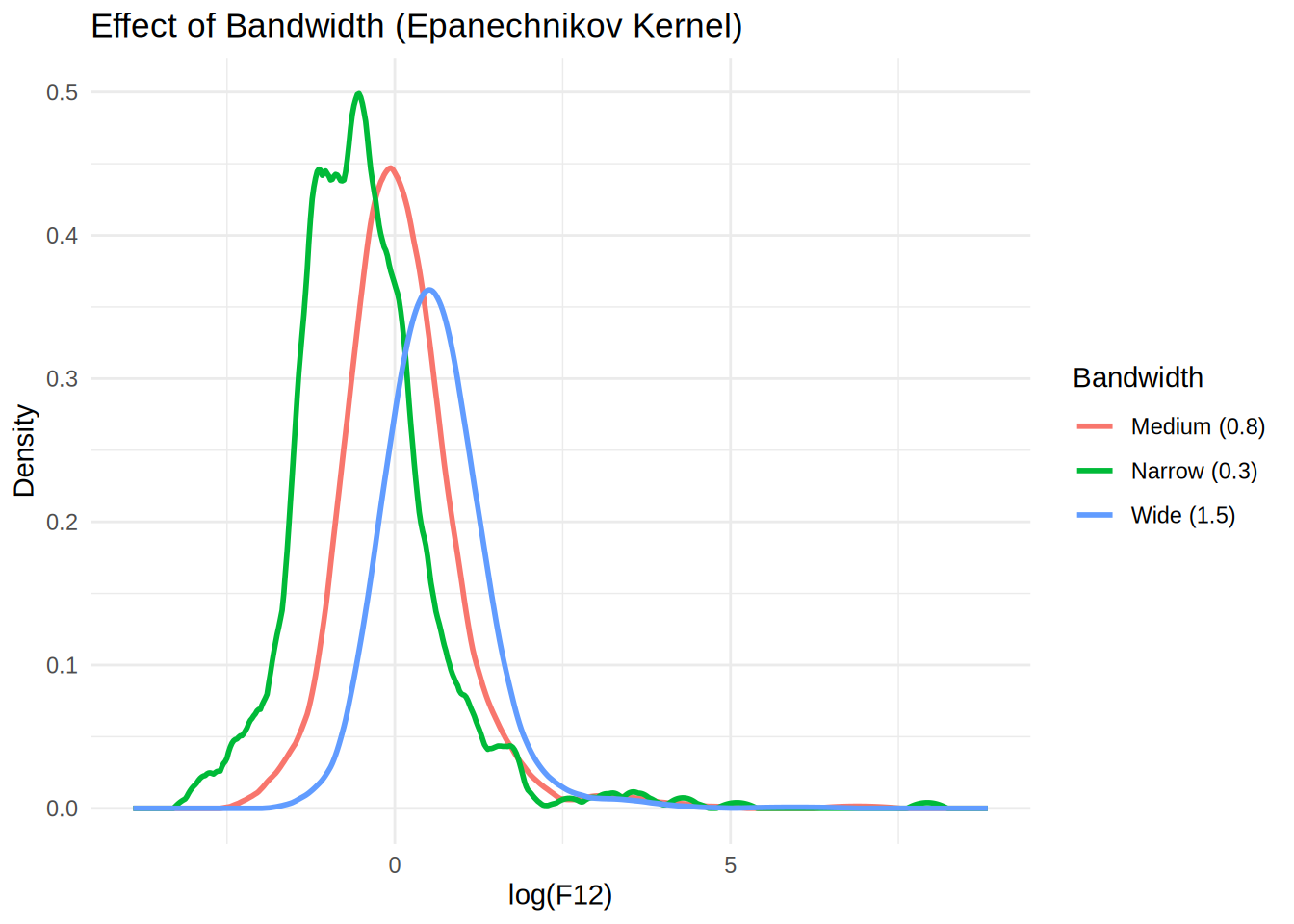
print(p2)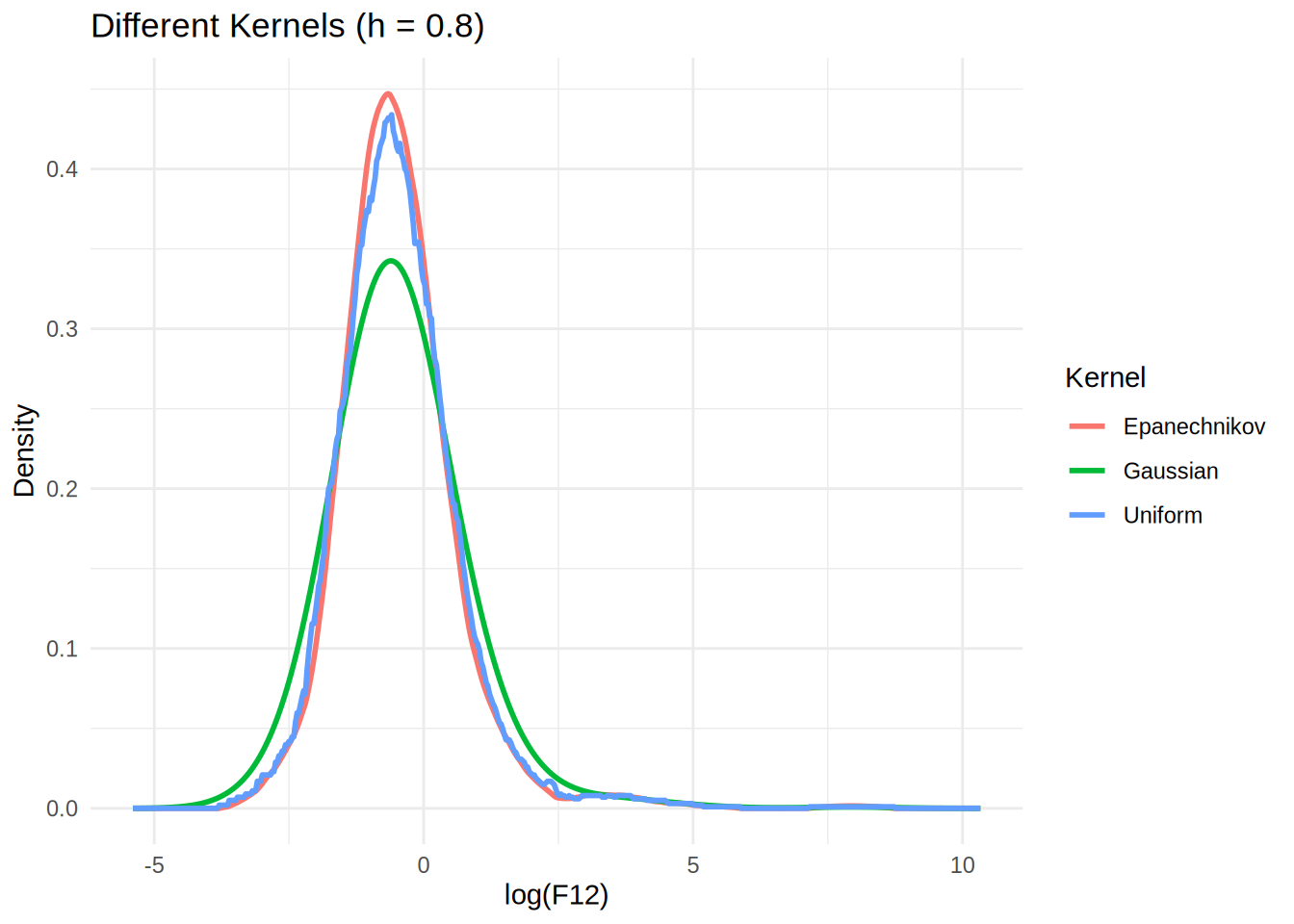
Key Benefits of This Currying Approach:
- Modularity: Each component (kernel, bandwidth) can be modified independently
- Reusability: Once created, curried functions can be reused multiple times
- Readability: Code becomes more self-documenting (e.g.,
epa_narrowclearly indicates both kernel and bandwidth) - Flexibility: Easy to create new combinations without rewriting core logic
- Functional Programming: Demonstrates higher-order functions and partial application
Advanced Example: Pipeline Processing
# Create a pipeline for comparing multiple bandwidth settings
bandwidth_comparison <- function(data, bandwidths) {
kde_functions <- lapply(bandwidths, function(bw) {
make_bandwidth_kde(epa_kde, bw)
})
results <- lapply(kde_functions, function(f) f(data))
names(results) <- paste("bw", bandwidths, sep = "_")
return(results)
}
# Use the pipeline
comparison_results <- bandwidth_comparison(log_F12, c(0.2, 0.5, 1.0, 1.5))
# Each result is readily available
str(comparison_results$bw_0.5, max.level = 1)List of 2
$ x: num [1:512] -4.5 -4.47 -4.44 -4.41 -4.39 ...
$ y: num [1:512] 0 0 0 0 0 0 0 0 0 0 ...str(comparison_results$bw_1.0, max.level = 1) NULLstr(comparison_results$bw_1.5, max.level = 1)List of 2
$ x: num [1:512] -7.5 -7.46 -7.42 -7.38 -7.34 ...
$ y: num [1:512] 0 0 0 0 0 0 0 0 0 0 ...This example shows how currying transforms our kernel density estimation from a monolithic function into a flexible, composable system. It’s the same principle we used with the .na.rm_decorator - we’re creating specialized functions from more general ones!